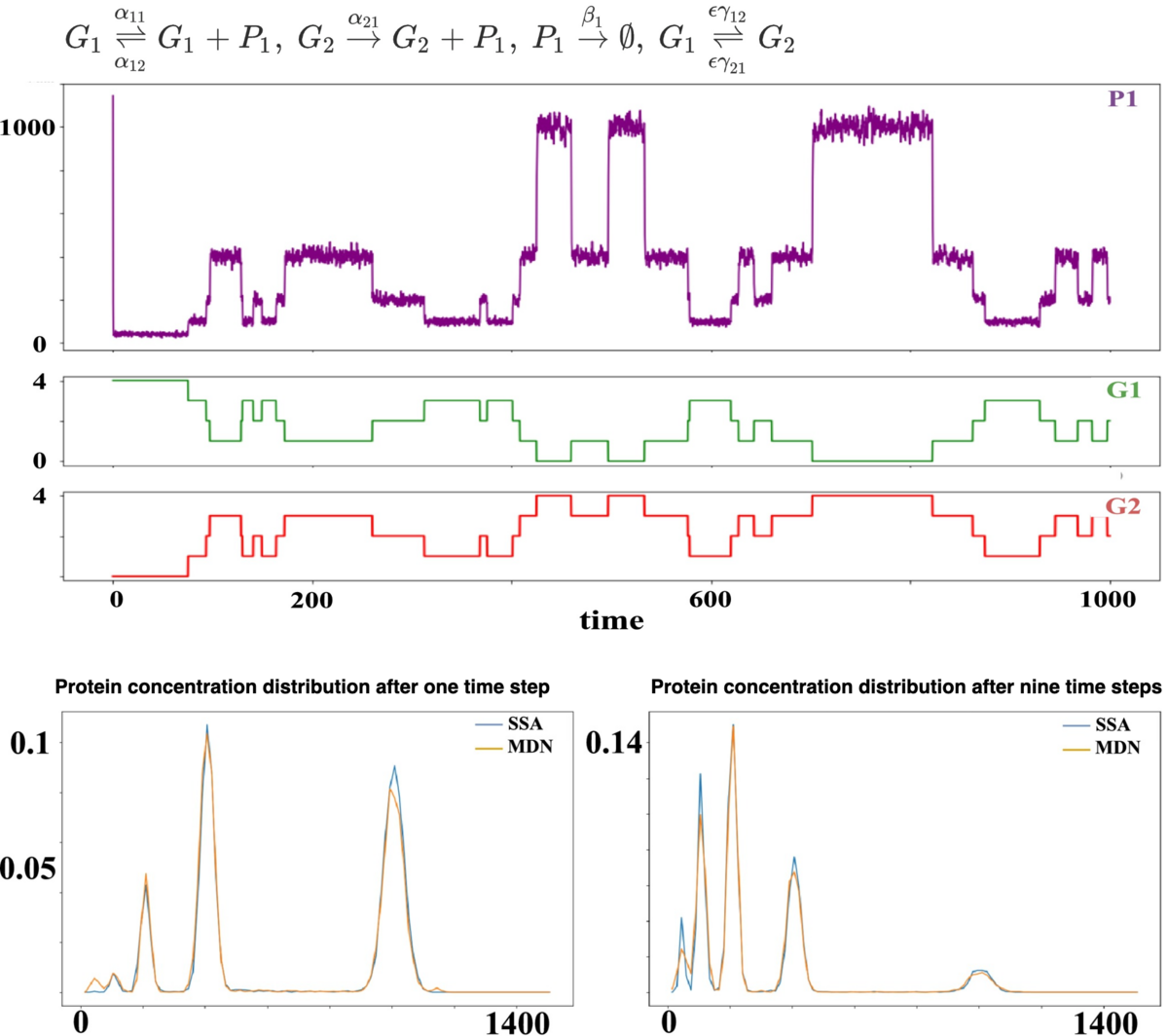
Automated reductions for stochastic Chemical Reaction Networks
Chemical reaction networks (CRNs) are used for modelling population dynamics in a wide range of sciences including organic chemistry, epidemiology, systems/synthetic biology, ecology. CRN models give rise to a continuous-time Markov chain (CTMC) which is often computationally demanding and even prohibitive to analyse in practice. The challenge is especially prominent in systems and synthetic biology research, where one deals with a large number of species, stochasticity, and events happening at multiple time-scales. It is hence desirable to develop automated model reduction techniques that allow for efficient simulation yet faithfully abstract the context-relevant features emerging from the hypothesised mechanism.
Related works
Tatjana Petrov, Denis Repin: Automated Deep Abstractions for Stochastic Chemical Reaction Networks, Information and Computation, 2021 [doi, pdf]
Repin Denis, Phung N Huy, Petrov Tatjana: StochNetV2: A Tool for Automated Deep Abstractions for Stochastic Reaction Networks, Quantitative Evaluation of Systems (QEST 2020) [doi, pdf]
Andreea Beica, Jérôme Feret, Tatjana Petrov: Tropical Abstraction of Biochemical Reaction Networks with Guarantees,The Ninth International Workshop on Static Analysis in Systems Biology (SASB 2018) [doi, pdf, poster]
Andreea Beica, Calin C. Guet, Tatjana Petrov: Efficient Reduction of Kappa Models by Static Inspection of the Rule-Set, Lecture Notes in Computer Science, Revised Selected Papers from Hybrid Systems Biology (HSB 2015) [doi, pdf]
Arnab Ganguly*, Tatjana Petrov*, Heinz Koeppl: Markov chain aggregation and its applications to combinatorial reaction networks (authors marked * with equal contributions), Journal of Mathematical Biology (JMB), 2014 [doi pdf
Tatjana Petrov: Formal reductions of stochastic rule-based models of biochemical systems (PhD thesis), ETH Zurich, 2013 [doi]
Tatjana Petrov, Heinz Koeppl: Approximate reductions of rule-based models, In Proceedings of ECC (European Control Conference), 2013 [doi pdf]
Tatjana Petrov, Jerome Feret and Heinz Koeppl: Reconstructing species-based dynamics from reduced stochastic rule-based models, In Proceedings of Winter Simulation Conference (WSC), 2012 [doi pdf]
Tom Henzinger, Jerome Feret, Heinz Koeppl, Tatjana Petrov: Lumpability abstractions of Rule-based systems, Journal of Theoretical Computer Science, 2011 [doi pdf]
Michael Klann, Loic Pauleve, Tatjana Petrov, Heinz Koeppl: Coarse-Grained Brownian Dynamics Simulation of Rule-Based Models, in Proceedings of the 11th Computational Methods in Systems Biology (CMSB), 2013 [doi pdf slides]
Ferdinanda Camporesi, Jerome Feret, Heinz Koeppl, Tatjana Petrov: Combining model reductions, 26th Conference on the Mathematical Foundations of Programming Semantics - MFPS, 2010 [doi pdf]
